Christian Ledig
Quick Links
Research Domain
I have substantial expertise in machine learning, computer vision and medical image analysis.Within a team at Twitter, UK I worked on, for example, “Photo-Realistic Single Image Super-Resolution Using a Generative Adversarial Network”. The paper is available here: [SRGAN]
During my PhD and PostDoc at Imperial College London, my general research domain was medical image processing with particular focus on the analysis and biomarker extraction of potentially abnormal magnetic resonance (MR) brain scans (e.g. [MALPEM], [DeepMedic]).
I am particularly interested in:
- machine learning and classification (e.g. random forests, manifold learning, deep learning)
- generative modelling (e.g. GANs, GMM, latent trees)
- domain adaptation / transfer learning
- image and video super-resolution
- classification of incomplete data / outlier prediction
- temporal modelling of disease progression
- image registration
- development and evaluation of image segmentation algorithms
- prediction and diagnosis support for dementia (e.g. AD) and traumatic brain injury (TBI)
Selected research topics
Photo-Realistic Single Image Super-Resolution Using a Generative Adversarial Network
Despite the breakthroughs in accuracy and speed of single image super-resolution using faster and deeper convolutional neural networks, one central problem remains largely unsolved: how do we recover the finer texture details when we super-resolve at large upscaling factors? The behavior of optimization-based super-resolution methods is principally driven by the choice of the objective function. Recent work has largely focused on minimizing the mean squared reconstruction error. The resulting estimates have high peak signal-to-noise ratios, but they are often lacking high-frequency details and are perceptually unsatisfying in the sense that they fail to match the fidelity expected at the higher resolution. In this paper, we present SRGAN, a generative adversarial network (GAN) for image super-resolution (SR). To our knowledge, it is the first framework capable of inferring photo-realistic natural images for 4x upscaling factors. To achieve this, we propose a perceptual loss function which consists of an adversarial loss and a content loss. The adversarial loss pushes our solution to the natural image manifold using a discriminator network that is trained to differentiate between the super-resolved images and original photo-realistic images. In addition, we use a content loss motivated by perceptual similarity instead of similarity in pixel space. Our deep residual network is able to recover photo-realistic textures from heavily downsampled images on public benchmarks. An extensive mean-opinion-score (MOS) test shows hugely significant gains in perceptual quality using SRGAN. The MOS scores obtained with SRGAN are closer to those of the original high-resolution images than to those obtained with any state-of-the-art method. C. Ledig, L. Theis, F. Huszar, J. Caballero, A. Cunningham, A. Acosta, A. Aitken, A. Tejani, J. Totz, Z. Wang, W. Shi, “Photo-Realistic Single Image Super-Resolution Using a Generative Adversarial Network”, accepted at CVPR (oral), 2017.
[pdf]
[bib]
C. Ledig, L. Theis, F. Huszar, J. Caballero, A. Cunningham, A. Acosta, A. Aitken, A. Tejani, J. Totz, Z. Wang, W. Shi, “Photo-Realistic Single Image Super-Resolution Using a Generative Adversarial Network”, accepted at CVPR (oral), 2017.
[pdf]
[bib]
Differential Dementia Diagnosis on Incomplete Data with Latent Trees
This software is also available for [download].Incomplete patient data is a substantial problem that is not sufficiently addressed in current clinical research. Many published methods assume both completeness and validity of study data. However, this assumption is often violated as individual features might be unavailable due to missing patient examination or distorted/wrong due to inaccurate measurements or human error. In this work we propose to use the Latent Tree (LT) generative model to address current limitations due to missing data. We show on 491 subjects of a challenging dementia dataset that LT feature estimation is more robust towards incomplete data as compared to mean or Gaussian Mixture Model imputation and has a synergistic effect when combined with common classifiers (we use SVM as example). We show that LTs allow the inclusion of incomplete samples into classifier training. Using LTs, we obtain a balanced accuracy of 62% for the classification of all patients into five distinct dementia types even though 20% of the features are missing in both training and testing data (68% on complete data). Further, we confirm the potential of LTs to detect outlier samples within the dataset.
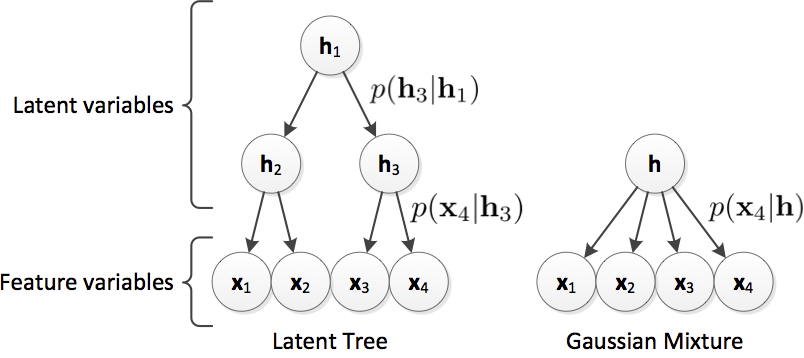 C. Ledig, S. Kaltwang, A. Tolonen, J. Koikkalainen, P. Scheltens, F. Barkhof, H. Rhodius-Meester, B. Tijms, A. W. Lemstra, W. Van der Flier, J. Loetjoenen, D. Rueckert, “Differential Dementia Diagnosis on Incomplete Data with Latent Trees”, Accepted in Medical Image Computing and Computer-Assisted Intervention MICCAI, 2016.
[pdf]
[bib]
[download]
C. Ledig, S. Kaltwang, A. Tolonen, J. Koikkalainen, P. Scheltens, F. Barkhof, H. Rhodius-Meester, B. Tijms, A. W. Lemstra, W. Van der Flier, J. Loetjoenen, D. Rueckert, “Differential Dementia Diagnosis on Incomplete Data with Latent Trees”, Accepted in Medical Image Computing and Computer-Assisted Intervention MICCAI, 2016.
[pdf]
[bib]
[download]
MALP-EM: Robust whole-brain segmentation of magnetic resonance images
This software is also available for [download].In recent years, multi-atlas segmentation has emerged as one of the most accurate techniques for the segmentation of brain magnetic resonance (MR) images, especially when combined with intensity-based refinement techniques such as graph-cut or expectation-maximization (EM) optimization. However, most of the work so far has focused on intensity-based refinement strategies for individual anatomical structures such as the hippocampus. In this work we extend a previously proposed framework for labeling whole brain scans by incorporating a global and stationary Markov random field that ensures the consistency of the neighbourhood relations between structures with an a-priori defined model. This flexible framework was successfully applied to automatically label brain MR scans with significant pathology present of patients suffering both dementia such as Alzheimer's disease or traumatic brain injury.
MALP-EM was evaluated as a Top 3 method in a Grand Challenge on whole-brain segmentation at MICCAI 2012.
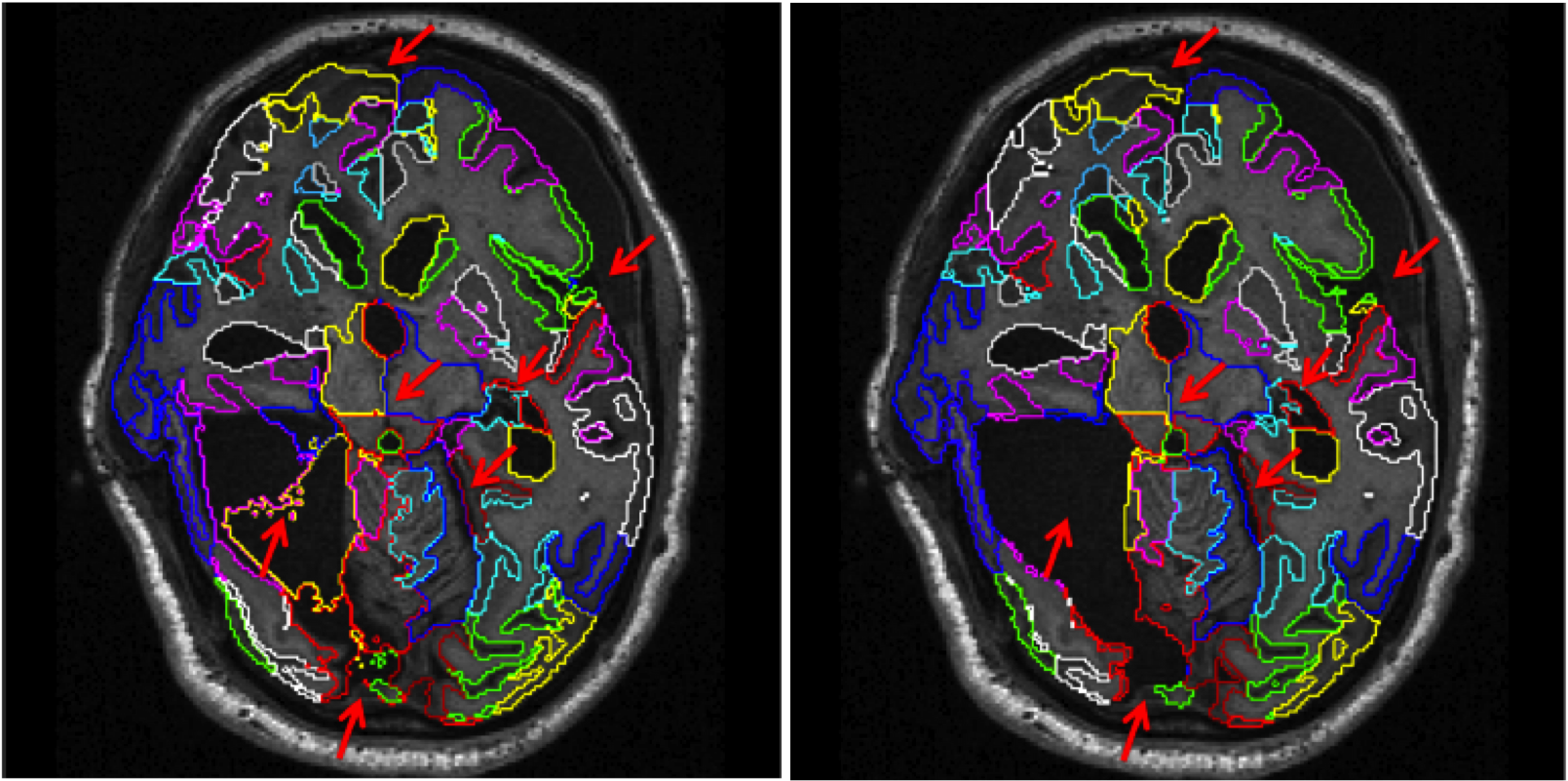 C. Ledig, R. A. Heckemann, A. Hammers, J. C. Lopez, V. F. J. Newcombe, A. Makropoulos, J. Loejoenen, D. Menon and D. Rueckert, “Robust whole-brain segmentation: application to traumatic brain injury,” Medical Image Analysis, 2015, in press. [doi]
C. Ledig, R. A. Heckemann, A. Hammers, J. C. Lopez, V. F. J. Newcombe, A. Makropoulos, J. Loejoenen, D. Menon and D. Rueckert, “Robust whole-brain segmentation: application to traumatic brain injury,” Medical Image Analysis, 2015, in press. [doi]
C. Ledig, R. A. Heckemann, P. Aljabar, R. Wolz, J. V. Hajnal, A. Hammers, and D. Rueckert, “Segmentation of MRI brain scans using MALP-EM,” MICCAI 2012 Grand Challenge and Workshop on Multi-Atlas Labeling, pp. 79-82, 2012. [pdf] [bib]
C. Ledig, R. Wolz, P. Aljabar, J. Loetjoenen, R. A. Heckemann, A. Hammers, and D. Rueckert, “Multi-class brain segmentation using atlas propagation and EM-based refinement,” Proceedings of ISBI 2012, pp. 896-899, 2012. [pdf] [bib] [doi]
Consistent longitudinal image segmentation
We propose a consistent approach to automatically segmenting longitudinal magnetic resonance scans of pathological brains. Using symmetric intra-subject registration, we align corresponding scans. In an expectation-maximization framework we exploit the availability of probabilistic segmentation estimates to perform a symmetric intensity normalisation. We introduce a novel technique to perform symmetric differential bias correction for images in presence of pathologies. To achieve a consistent multi-time-point segmentation, we propose a patch-based coupling term using a spatially and temporally varying Markov random field. We demonstrate the superior consistency of our method by segmenting repeat scans into 134 regions. Furthermore, the approach has been applied to segment baseline and six month follow-up scans from 56 patients who have sustained traumatic brain injury (TBI). We find significant correlations between regional atrophy rates and clinical outcome: Patients with poor outcome showed a much higher thalamic atrophy rate (4.9+/-3.4%) than patients with favourable outcome (0.6+/-1.9%).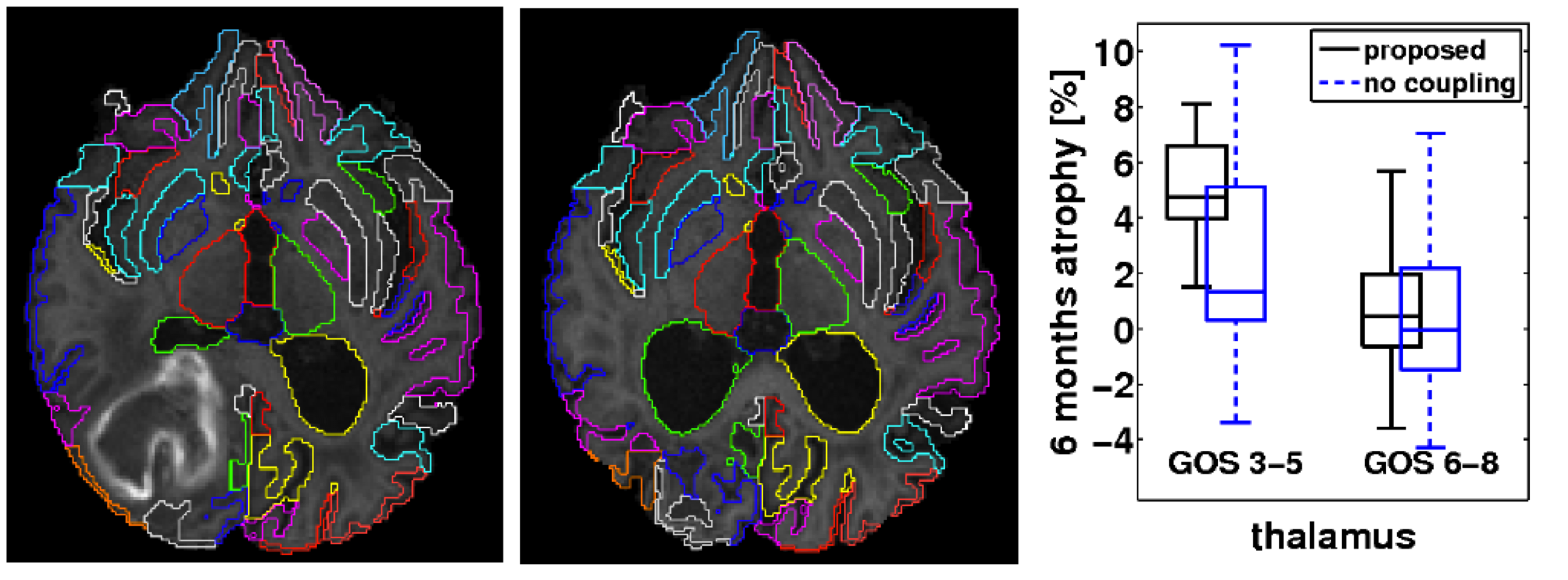 C. Ledig, W. Shi, A. Makropoulos, J. Koikkalainen, R. A. Heckemann, A. Hammers, J. Loetjoenen, and D. Rueckert, “Consistent and robust 4D whole-brain segmentation: application to traumatic brain injury,” accepted at ISBI, 2014.
[pdf]
[bib]
C. Ledig, W. Shi, A. Makropoulos, J. Koikkalainen, R. A. Heckemann, A. Hammers, J. Loetjoenen, and D. Rueckert, “Consistent and robust 4D whole-brain segmentation: application to traumatic brain injury,” accepted at ISBI, 2014.
[pdf]
[bib]C. Ledig, V. Newcombe, M. G. Abate, J. G. Outtrim, D. Chatfield, T. Geeraerts, A. Manktelow, P. J. Hutchinson, J. P. Coles, G.Williams, D. Rueckert, and D. Menon, “Dynamic evolution of atrophy after traumatic brain injury,” Abstract accepted at ISMRM, 2014. [bib]
Neonatal tissue segmentation
The dynamic contrast changes in T2-weighted magnetic resonance (MR) images in developing brains are one of the main reasons that make tissue segmentation in neonatal MR brain scans very challenging. However, to assess and understand early brain development, it is crucial to be table to robustly and accurately segment neonatal MR images. It is especially the varying intensity profile from white matter (WM) to grey matter (GM) to cerebrospinal uid (CSF) that hampers the direct application of intensity based segmentation algorithms such as the expectation-maximization (EM) algorithm. In this work, we present an approach that incorporates prior spatial information by using a 4D probabilistic spatio-temporal atlas and extends the widely used EM algorithm by an energy function that penalizes implausible neighborhood configurations. Inspired by the Markov Random Field (MRF) that is described by a connectivity matrix, we introduce a connectivity tensor that allows the incorporation of second order neighborhood information. We evaluate our approach by careful visual inspection and by a measure based on the number and size of connected WM components.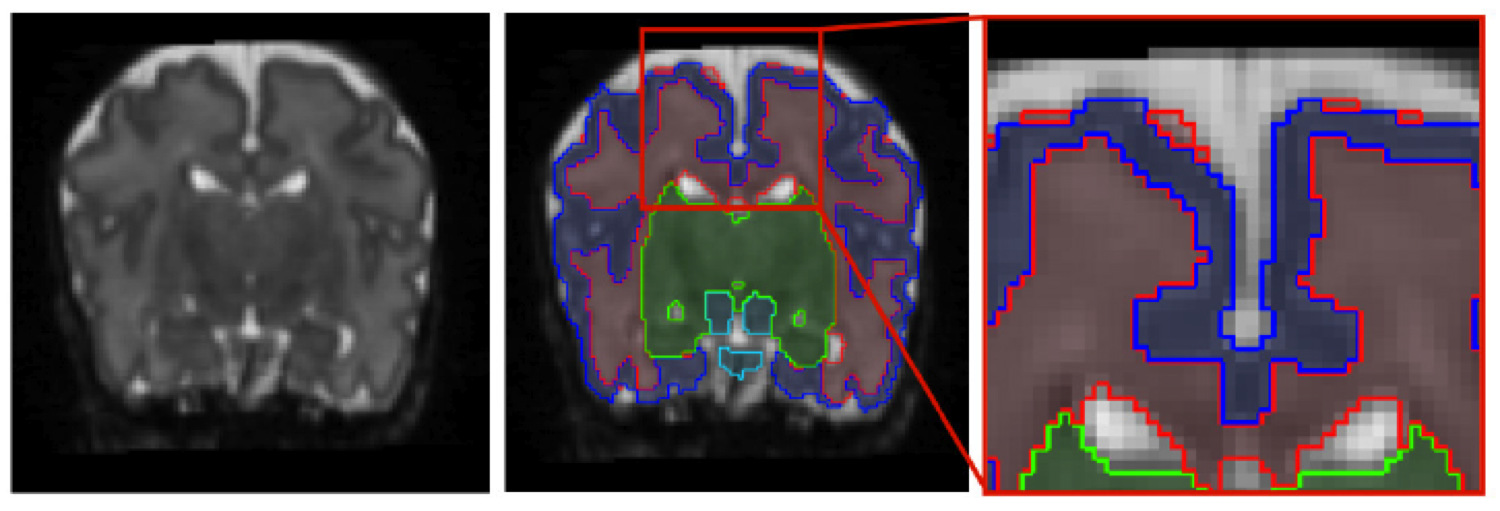 C. Ledig, R. Wright, A. Serag, P. Aljabar, and D. Rueckert,
“Neonatal brain segmentation using second order neighbourhood
information,” MICCAI PAPI, pp. 33-40, 2012.
[pdf]
[bib]
C. Ledig, R. Wright, A. Serag, P. Aljabar, and D. Rueckert,
“Neonatal brain segmentation using second order neighbourhood
information,” MICCAI PAPI, pp. 33-40, 2012.
[pdf]
[bib]R. Wright, V. Kyriakopoulou, C. Ledig, M. Rutherford, J. V. Hajnal, D. Rueckert, and P. Aljabar, “Automatic quantification of normal cortical folding patterns from foetal brain MRI,” NeuroImage, 2014. [bib] [doi]
Evaluation of image segmentation
This software is also available for [download].The quantification of similarity between image segmentations is a complex yet important task. The ideal similarity measure should be unbiased to segmentations of different volume and complexity, and be able to quantify and visualise segmentation bias. Similarity measures based on overlap, e.g. Dice score, or surface distances, e.g. Hausdorff distance, clearly do not satisfy all of these properties. To address this problem, we introduce Patch-based Evaluation of Image Segmentation (PEIS), a general method to assess segmentation quality. Our method is based on finding patch correspondences and the associated patch displacements, which allow the estimation of segmentation bias. We quantify both the agreement of the segmentation boundary and the conservation of the segmentation shape. We further assess the segmentation complexity within patches to weight the contribution of local segmentation similarity to the global score. We evaluate PEIS on both synthetic data and two medical imaging datasets. On synthetic segmentations of different shapes, we provide evidence that PEIS, in comparison to the Dice score, produces more comparable scores, has increased sensitivity and estimates segmentation bias accurately. On cardiac magnetic resonance (MR) images, we demonstrate that PEIS can evaluate the performance of a segmentation method independent of the size or complexity of the segmentation under consideration. On brain MR images, we compare five different automatic hippocampus segmentation techniques using PEIS. Finally, we visualise the segmentation bias on a selection of the cases.
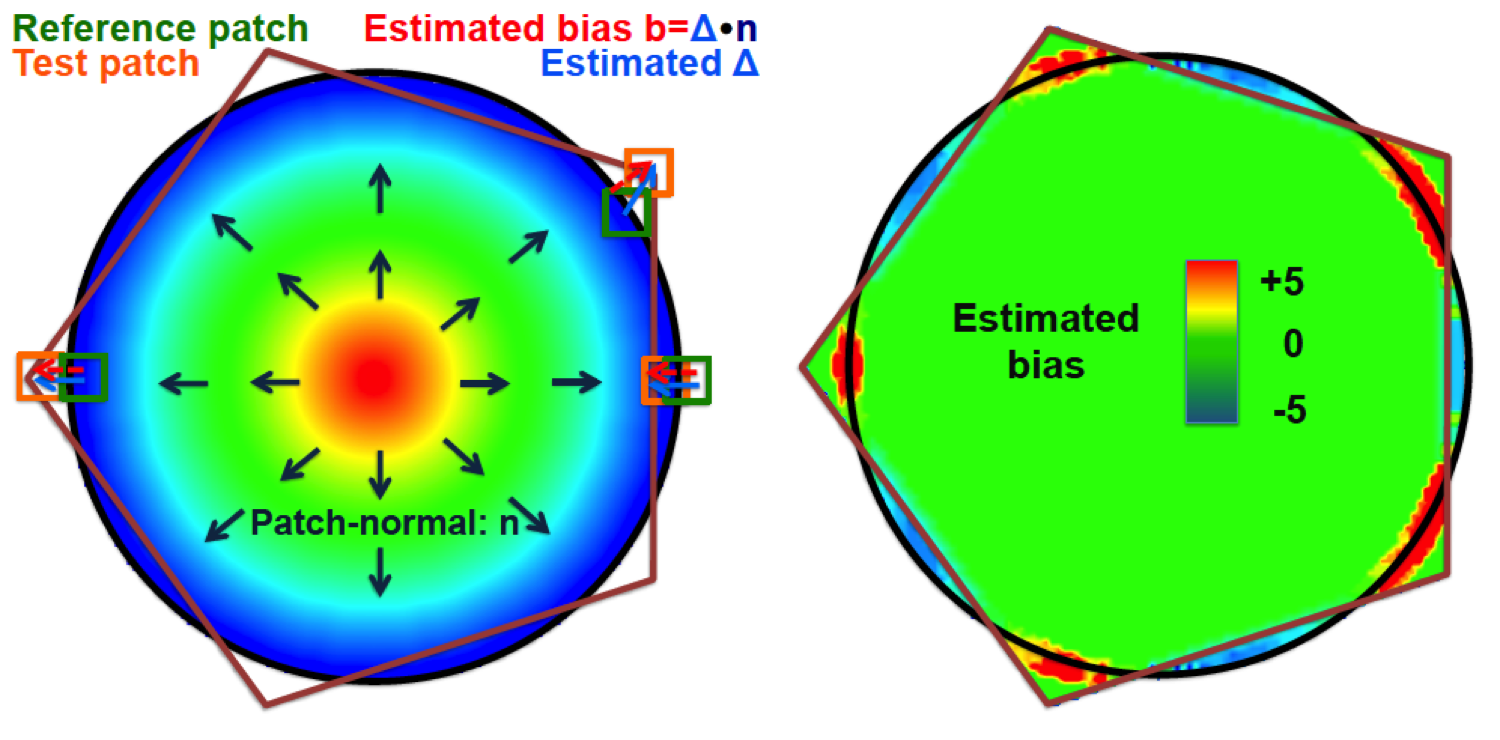 C. Ledig, W. Shi, W. Bai, and D. Rueckert, “Patch-based evaluation of image segmentation,” CVPR, pp. 3065-3072, 2014.
[bib]
[pdf]
[spotlight:mpeg4][spotlight:mov]
[download]
C. Ledig, W. Shi, W. Bai, and D. Rueckert, “Patch-based evaluation of image segmentation,” CVPR, pp. 3065-3072, 2014.
[bib]
[pdf]
[spotlight:mpeg4][spotlight:mov]
[download]
 @LedigChr
@LedigChr



